 
Ten mL of TNA DNA was used to generate cDNA. We extracted total nucleic acid (TNA) from 200 mL of each nasal swab using the EasyMag Extraction Platform (Biomerieux, Marcy l’Etoile, France). We employed MassTag polymerase chain reaction (PCR) to identify the presence of 22 common bacterial and viral respiratory pathogens (see Table S1). As previously described, swabs were stored at −80 ☌ until analysis all samples were shipped from Ghana to the United States on dry ice with internal temperature monitors. These analyses include only children with no respiratory symptoms at time of swab and who did not have a physician-diagnosed pneumonia episode within two weeks prior to or following swab collection. Nasopharyngeal Microbiota Sampling and PCRĪs above, NPM swabs were collected in infants with and without respiratory symptoms. The current analyses include a convenience sample of N = 112 children without respiratory symptoms at the time of nasopharyngeal swab and who successfully performed IOS at child age four years. Children are being followed in ongoing prospective work with respiratory phenotyping, including lung function measured by IOS at child age four years. Nasopharyngeal microbiota (NPM) swabs were collected for children with physician-diagnosed pneumonia and age- and sex-matched healthy controls (total N = 669). Fieldworkers trained in the World Health Organization’s Integrated Management of Childhood Illness (IMCI) performed weekly surveillance of study children over the first year of life and referred any child meeting criteria for pneumonia or who was otherwise unwell for physician evaluation and pneumonia diagnosis. Key individual and household level data were collected on enrollment, including maternal age, parity, ethnicity, and number of people living in the household. 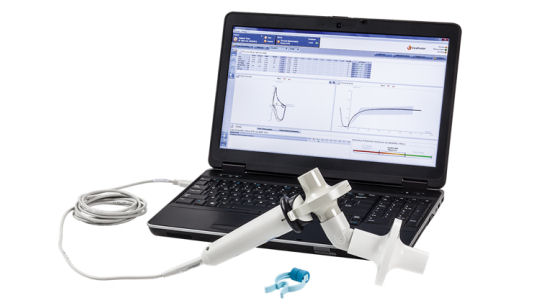
Briefly, pregnant nonsmoking women were recruited prior to ultrasound-confirmed 24 weeks of gestation. The GRAPHS cohort has been described elsewhere. Mother–infant dyads were recruited from a pregnancy cohort derived from the Ghana Randomized Air Pollution and Health Study (GRAPHS) with ongoing, prospective follow up through child age four years. It is unknown whether pathogen colonization is linked to other objective measures of future lung health, such as lung function. Data from our cohort in rural Ghana suggests a possible link between early life colonization with bacterial pathogens, specifically Streptococcus pneumonia, Haemophilus influenza, Moraxella cattarhalis, and infant pneumonia. However, when they appear earlier, they are often associated with increased risk of wheeze and the development of asthma. Streptococcus, Haemophilus, and Moraxella species tend to colonize the respiratory tract later in development. Specifically, Cornebacteriacae and Staphylococcus species which are earlier colonizers of the respiratory tract, may decrease risk for childhood wheeze and asthma. Disruptions in microbiome maturation may lead to airway inflammation, chronic wheeze and asthma, and pneumonia. Further studies are required to understand the role of the microbiota in future lung health.ĭysbiosis, or alterations in microbe colonization of the upper airway, has been associated with respiratory disease. In multivariable linear regression models, the less diverse NPM subphenotype had higher small airway resistance (R5-R20 β = 17.9%, 95% CI 35.6, 0.23 p = 0.047) compared with the more diverse subphenotype. We identified a higher diversity subphenotype (N = 38, 34%) with more pathogens (median 4 IQR 3.25, 4.75) and a lower diversity subphenotype (N = 74, 66%) with fewer pathogens (median 1 IQR 1, 2). LCA suggest that a two-class model best described the infant NPM. Then, we used linear regression to analyze associations between subphenotype assignment and lung function. First, we employed latent class analysis (LCA) to identify nasopharyngeal microbiota (NPM) subphenotypes. We prospectively followed the cohort and measured lung function at age four years by impulse oscillometry. Leveraging the Ghana Randomized Air Pollution and Health Study (GRAPHS) prospective pregnancy cohort in Kintampo, Ghana, we collected nasopharyngeal swabs in 112 asymptomatic children aged median 4.3 months (interquartile range (IQR) 2.9, 7.1) and analyzed 22 common bacterial and viral pathogens with MassTag polymerase chain reaction (PCR). 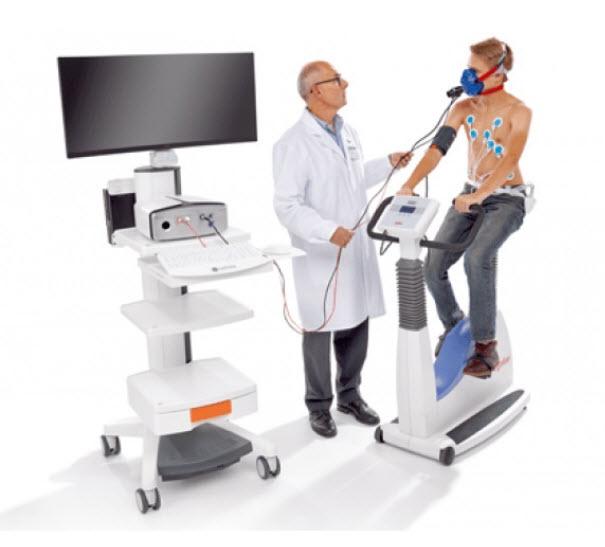
Associations between respiratory microbiota and lung health in children from low- and middle-income countries are not well-described. Early life respiratory microbiota may increase risk for future pulmonary disease. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
